A new software tool makes it easier to study relationships between a host, its microbiome and pathogens like HIV or SARS-CoV-2.
SAN ANTONIO (May 4, 2022) – Researchers at Texas Biomedical Research Institute and Tulane University have developed a new software tool that makes it easier, faster and more cost effective to analyze genetic information about a host and its microbiome at the same time. The software, called “meta-transcriptome detector” (MTD), can be used by a wide range of microbiologists and drug developers, including those researching diseases such as certain cancers, COVID-19, HIV/AIDS, malaria and many other human health conditions linked to microorganisms. The tool was recently published in the journal Briefings in Bioinformatics.

“It is very user-friendly, especially for researchers with little to no background in bioinformatics,” says Associate Professor Binhua “Julie” Ling, PhD, who co-leads Texas Biomed’s Host-Pathogen Interactions Research Program and is the senior author of the paper. “You only need to write one line of code to set certain parameters and the software does the rest automatically.”
MTD enables researchers to get a comprehensive snapshot of what microbes are present in the host – including the huge range of “good bacteria” that normally live on and inside people and animals, as well as the harmful ones, such as viruses that cause serious illnesses. More importantly, MTD analyzes gene expression activity, essentially which genes are turned on or off, in both the microbes and the host simultaneously, allowing researchers to easily spot relationships between them. For example, seeing which genes are active in a microbe and in the host could indicate that the activity of one is influenced by the other, and might suggest a potential target for treatment down the line, Ling explains.
“We can’t say at this stage if it is cause and effect, but we can use this analysis to pinpoint what genes or pathways we should be investigating – perhaps ones that we never considered before as being related,” Ling says. “MTD can help accelerate that process and potentially open new avenues of research and drug development.”
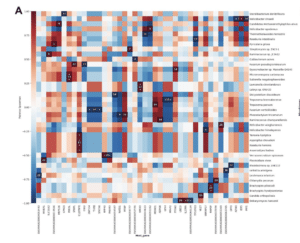
MTD links together several existing software packages and pulls from international databases containing RNA sequences of more than 100,000 species of bacteria, viruses, fungi, archaea and protozoa, as well as sequences of vectors and plasmids. Users are also able to update the database with specific sequences they are interested in.
First paper author Fei Wu, Ph.D., worked with Ling to study how the microbiome changes with age in younger and older monkeys with simian immunodeficiency virus (SIV), the monkey version of HIV.
“We were having to analyze gene expression from the host by one workflow, and the microbiome gene expression by a separate workflow,” Wu says. “We wondered why can’t we do both at the same time?”
Since their lab was relocating during the COVID-19 pandemic, they had the time to focus on computer-based work, and set out to solve this issue. Wu worked with Ling and collaborator Yao-Zhong Liu, PhD, an Associate Professor at Tulane University School of Public Health and Tropical Medicine, to build and test the new software.
“Normally, we are using bioinformatics software to analyze our data, not building it,” Wu says. “It was challenging, but exciting, to branch out and now have something that will not only help us, but also any other researcher doing RNA sequencing of hosts and microbiomes; from humans and monkeys, to mosquitos carrying malaria parasites and snails carrying schistosome parasites.”
MTD offers several advantages over existing tools. Primarily, it has the ability to analyze RNA sequences of both the host and the microbiota from the same sample – either single cells or a bulk tissue sample. This saves both time and resources, in addition to providing new insights into the relationships between the microorganisms and host. It does require some high-performance computing power, but otherwise is very accessible to researchers without much bioinformatics experience. For more information: read the paper, and check out the program and user guide on github.
Funding acknowledgement:
This work was supported by the National Institutes of Health Award Numbers R01 MH116844, R01 NS104016, and P51OD011104 and P51OD011133. The funders had no role in study design, data collection and analysis, preparation of the manuscript or decision for publication.
###
ABOUT TEXAS BIOMED
Texas Biomed is one of the world’s leading independent biomedical research institutions dedicated to eradicating infection and advancing health worldwide through innovative biomedical research. Texas Biomed partners with researchers and institutions around the world to develop vaccines and therapeutics against viral pathogens causing AIDS, hepatitis, hemorrhagic fever, tuberculosis and parasitic diseases responsible for malaria and schistosomiasis disease. The Institute has programs in host-pathogen interactions, disease intervention and prevention, and population health to understand the links between infectious diseases and other diseases such as aging, cardiovascular disease, diabetes and obesity. For more information on Texas Biomed, go to www.TxBiomed.org.