Research Focus
Dr. Ebrahimi leads a cross-disciplinary quantitative biology program. His lab combines data and expertise across multiple quantitative and experimental science disciplines such a genetics, virology, cancer, evolution, bioinformatics, mathematics, and statistics to develop and test novel hypotheses. Current studies in Dr. Ebrahimi’s lab focus on identification and quantification of molecular processes in viral and cancer immunity and evolution.
- Discover population-specific virus-host interaction mechanisms – Why do some immune genes (e.g. APOBEC3 enzymes) express and splice differently in different human populations? How do these differences play a role in the interaction between immune system and viruses such as HIV-1?
- Delineate genetic and mechanistic basis of antiviral and cancer immunity – Why do the rate and severity of infectious diseases and cancer vary in different humans? What are the viral and host contributing factors?
- Develop quantitative biology methods and tools to deconvolute complex biological processes – What are the sources of cancer mutations and gene expression variation in cancer? How many dysregulated processes are present in a given tumor tissue biopsy? Is viral infection a player in cancer initiation and/or evolution?
Dr. Ebrahimi has more than a decade of experience working in quantitative biology and genomics and uses multi-omic analysis and systems biology to answer questions such as those listed above.
Announcement
Research positions (postdoc, scientist, technicians, etc.) in quantitative biology are available. Qualified candidates with a background in bioinformatics or with a background in quantitative sciences such as statistics, mathematics, physics, engineering, and with a passion for learning medicine and biology and working on projects related to infectious diseases, cancer, and evolution, please email Dr. Ebrahimi a CV.
Inside the Lab
Infectious Diseases are a major cause of death and a significant socioeconomic burden on healthcare systems. According to UNAIDS, over 36 million people including 1.8 million children under 15 yrs old were infected by HIV in 2017. The rate of new infection worldwide was estimated to be around 5,000 per day, and most of the new infections were in sub-Saharan Africa. HIV infection and its associated illnesses such as tuberculosis, lymphomas, Kaposi sarcoma, and neurocognitive disorder are complex diseases. The immune processes involved in these diseases are numerous and many are yet to be identified and fully characterized. In my lab, data from various sources such as the genome, transcriptome, and proteome of patients and viruses such as HIV are analyzed to determine what viral and host immune factors play a role in our immunity against viruses. Our multi-omic analysis has revealed that some immune genes such as APOBEC3 undergo differential splicing in different human populations, and this may be one of the reasons for our differential immunity against viruses. We are using a combined bioinformatic/experimental approach to investigate the genetic basis and functional consequences of population-specific immune gene splicing and virus-host interaction mechanisms.

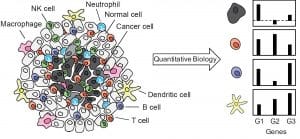
Cancer is a major health challenge and is estimated to be responsible for more than 600,000 deaths in the US in 2019. New diagnosed cancer cases is estimated to be greater than 1,700,000. There are over 100 different types of cancers, each with their own unique characteristics and many having similar features such as mutation profiles. Studies have shown that cancer can be caused by both internal and external factors. For example, viral infection or exposure to UV radiation are external cancer inducer factors. However, internal factors such as inherited mutations within DNA repair enzymes or methylation-induced mutations can also be a source of tumor initiation and/or evolution. Some of these factors are associated with distinct mutation, expression, and immune signatures in tumor tissue biopsies. Deconvolution of these molecular and cellular signatures is not trivial, because many different cells in the tumor and its microenvironment are in dynamic interactions with each other. We are developing combined quantitative biology/single cell analysis techniques to quantify the contribution of each cell type and each molecular process and determine their roles in cancer evolution and survival outcomes.
Main Technologies and Methods Used
- Next generation DNA and RNA sequencing
- Computational analysis, Bioinformatics
- Single cell analysis
- Cell culture and viral infectivity assays
- Fluorescence-Activated Cell Sorting (FACS)
- Spectroscopy
- Mass spectrometry
